Project Description
Doctor’s medical prescriptions are difficult to understand for not only semi literate people but also learned ones. This is mainly due to cryptic medical abbreviations used by the medical people and also because of the complex vocabulary. So a system which can help the people understand the drug prescription prescribed to them and know more about the other parameters of the drug such as side-effects, usage etc will be quite useful.
Thereby the proposed system would automatically convert the complex cryptic doctor's prescription to layman text for quick and better understanding with added functionalities of preferred language and more information about the drug and the diagnosis.
Project Architecture
The architecture of the project is as follows
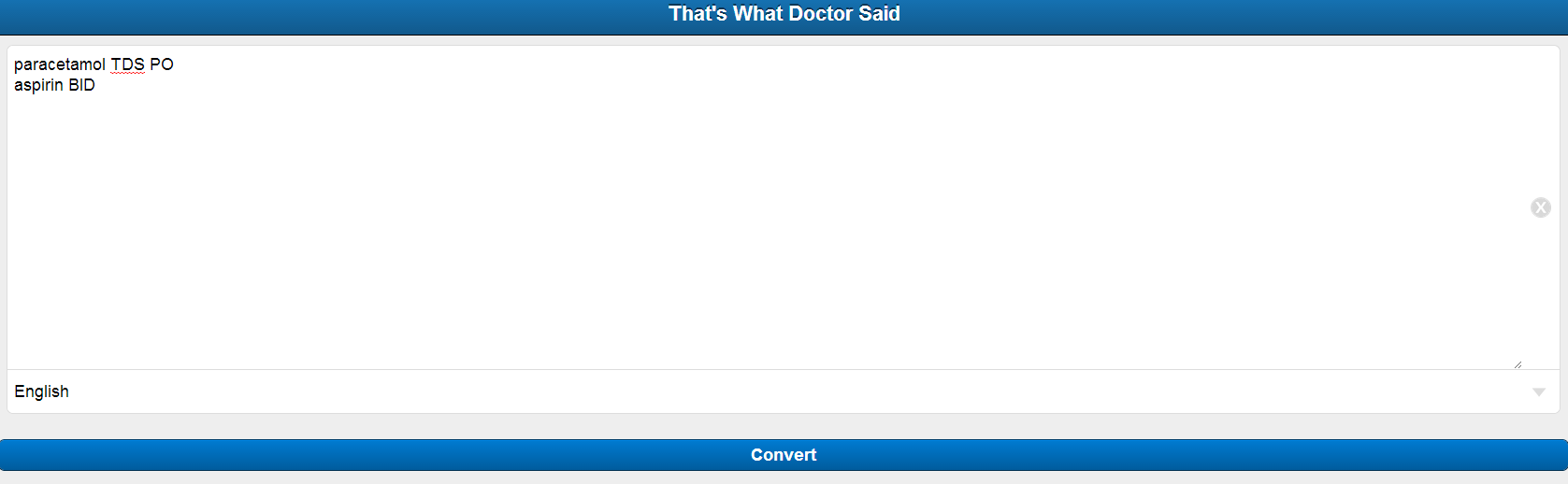
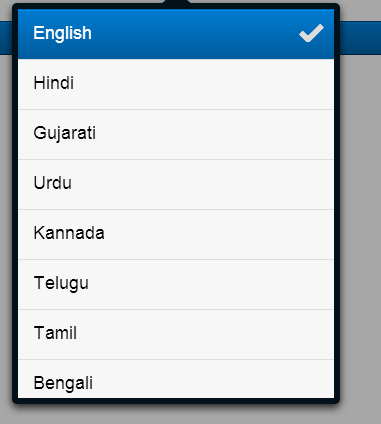
- A Sencha mobile application window will accept the user input and the preferred language of the output.
- The application will then send the query to the backend server via REST API call.
- Apache cTAKES backend server which will extract all the required entities such as drug/diagnosis name, frequency, strength, route, form etc from the user input.
- An exhaustive set of regular expressions will then extract the medical abbreviations and other entities not directly obtained from cTAKES.
- Then Natural language generator module will convert the form variables extracted from cTAKES into a natural text.
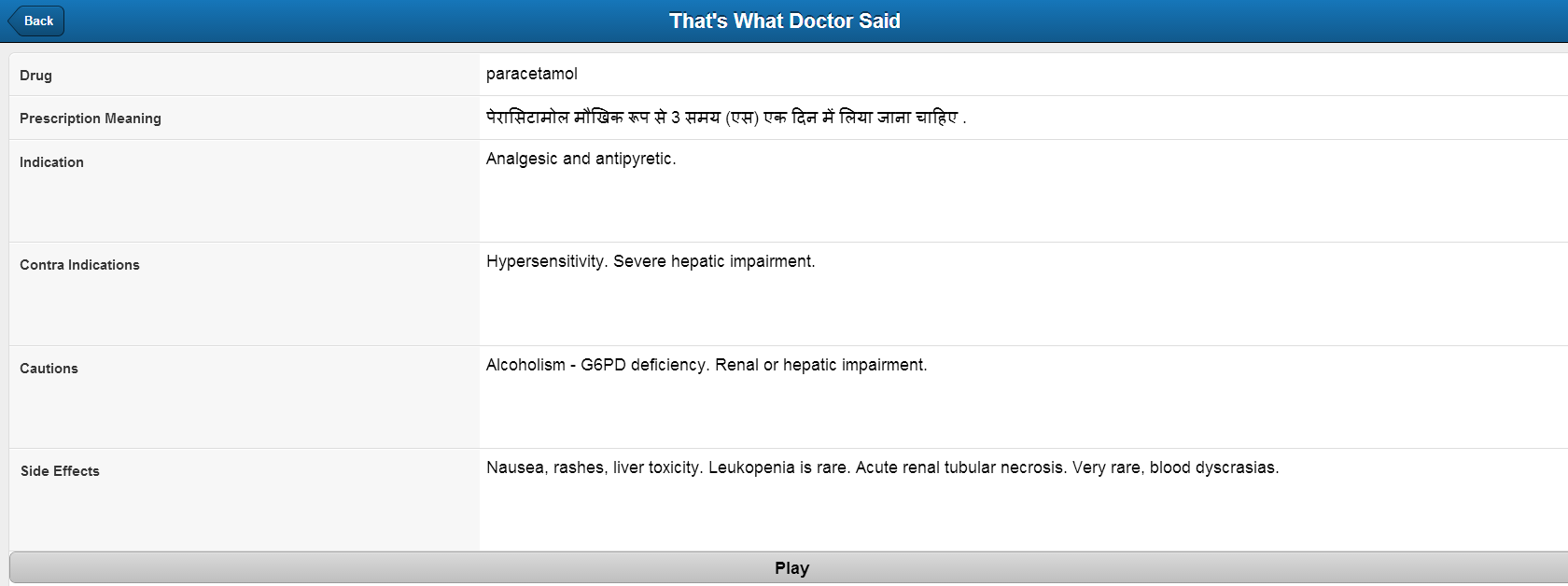
- Natural text is then translated to the preferred language of the user using Google API.
- Convert text is then sent back to the user application.
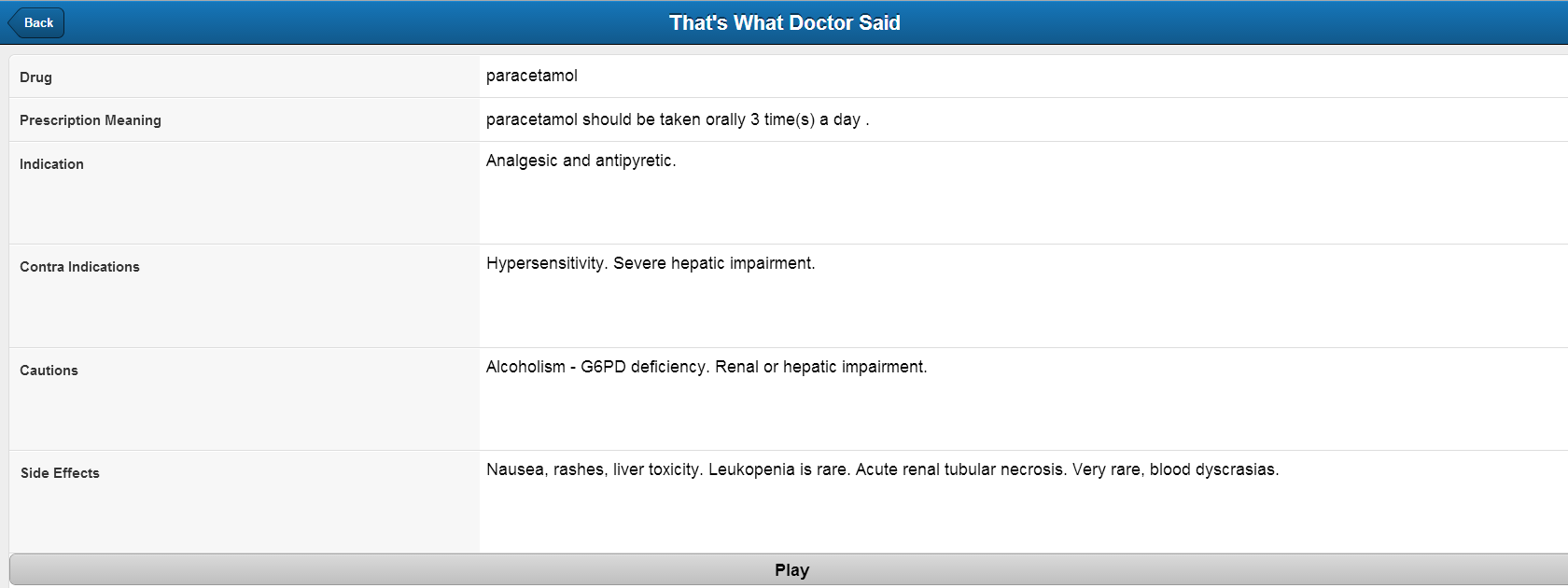
- For more information, the extracted drugs/diagnosis are looked upon on the WebMD and NLM website to find more information such as Uses, Side-effects, Warnings, Precautions etc.
Implementation Details
- Front end: Sencha
- Back end: Java, MySQL
- Libraries Used: Apache cTAKES, gtranslate API, JDBC MySQL driver
- Project Facets: Maven, Java Web Application
- Server: Tomcat 7
Projects Plan and Time-line with Milestones
May, 19 - May, 31 :
Goal : Complete the Front and back end for doctor module.
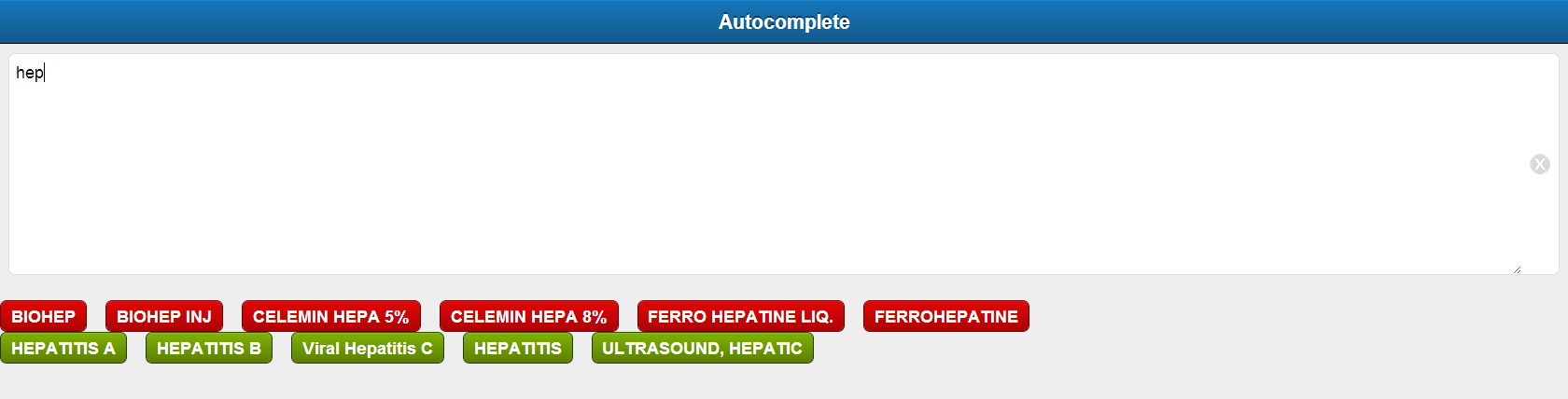
Make a Sencha web application for Raxa Doctor module. The application has a textbook with auto-complete feature. The applications listens for each letter typed in and uses it as a search key to get the relevant results from the backend server.
- Build a database of relevant observations and Complete the backend server (a search engine).
Integrate and test the module.
June, 1 - June, 15 :
Goal : Complete the cTakes Backend and start with Patient module front end.
Complete the cTAKES backend server to extract the required information from the text.
Make a list of regular expressions to extract more information from the input text so that accuracy of the Information Extraction module can be enhanced.
Build the layman text generator module to convert the information from the form variables into natural text.
Start with Raxa Patient module front end.
Integrate with Raxa Patient module <temp webapp> and test for accuracy of the extraction process.
June, 16 - June, 20 :
Goal : Complete the Language conversion task and test the whole module.
Integrate with the Google API to translate the text into the preferred language.
Build the Sencha mobile application to accept input and language preference from the user.
Complete the whole UI and test the application.
June, 21 - June 25 :
Goal : Midterm Evaluations
June, 26 - July, 05:
Goal : Complete the Information Retrieval (Drug/Diagnosis) service.
Devise ways to extract drug information from various websites such as WebMD, NLM, drugs.com etc.
Complete the module and test using REST API for various test cases.
July, 06 - July, 20 :
Goal : Complete the front end and Test for production
Prepare the front end for showing information about the drug/diagnosis.
Integrate with the Information Retrieval Component.
Complete the project and make necessary changes/addition to the UI.
Prepare SMS service for the same if time permits.
July, 21 - August, 15 :
Goal : Refractor, Clean and Document the code
August, 16 - August, 21 :
Goal : Demonstration and Documentation
August, 21 - August, 25
Goal : Final Evaluations
Detailed Description:
Patient Module
The project is a Maven project so the pom.xml files denotes all the libraries required by the project as dependencies. Other cTAKES resources required by the different annotators models are required to be downloaded manually though. We provide those with our project since we require only a subset of the resources.
- pom.xml
Dependencies- ctakes-drug-ner : For the ctakes library (only the drug extraction pipeline)
- jersey-core, jersey - server : For providing REST API to our ctakes - java based backend
- jersey - servlet : For holding the servlets on the server.
- jersey - json : To handle the JSON format required to format the output.
- gson: For conversion to JSON format
- mysql-connector-java: Driver to connect to MySQL database.
- External Resources
The folder SentenceDetection, Chunker, POSTagger and DictionaryLookup contains the external resources required by the ctakes server. The file LookupDesc_DrugNER.xml contains important configuration required to be filled before running the NER pipeline.
Gtanslate API (for language conversion)
- Java Sources
Package org.raxa contains the main project Java file while org.raxa.rest contains the sources related to providing the api.
- CtakesService.java
Is the main class which perform the core of the functions. It performs three main functions of a. Initializing the ctakes Analysis Engine and abbnList, b. Normalizing the input received c. Extracting the drug parameters using the initialised analysis engine and creating the Drug object. - NaturalLanguageGenerator.java
This class provides the method to convert to drug extracted in form of parameters mentioned above to free form natural text. As of now contains only one method getNaturalText() which takes Drug object as input and converts and gives the layman natural text as output. - Abbreviation.java
Helper class which provides abbreviation object to normalize the input text. It contains three parameters acronym, acronym-text, and acronym-type. Acronym is the short form (for eg TDS), acronymText is the full form (eg Three times a day) while acronymType is the category of acronym (drug-form, drug-frequency, drug-route) in our example drug-frequency. As of now these are three categories we divide the most frequently used acronyms into. These acronyms are read/inflated from the files separately provided in the Abbreviations folder. - Drug.java
Helper class which creates the Drug object for the drug extracted from the input. It contains the parametes drugName, drugFrequency, drugFrequencyUnit, drugStrength, drugStrengthUnit, drugRoute, drugForm, drugDuration and drugDosage. - InformationExtraction.java
This class provides the method to search for more information about the drug using Generic drug name. The information obtained are Indications, Contraindications, Precautions and Side effects.
- How to run?
Note : Will require Internet Connection to install as well as run the server- Plugins/Softwares Required
- Apache Maven
- Apache Tomcat Server
- Method 1.
- Download the Project and change working directory to the Patient folder.
- run mvn clean install
- This will generate a war file inside target directory.
- Place the war directory under the webapps folder of your tomcat installation.(/var/libs/tomcat7/webapps/)
- Use POSTMAN (Google Chrome app) to test working. The url is http://localhost:8080/ctakes/rest/ctakes/hello?text=Prescription&language=language
- Method 2. To run with eclipse
- Import the project.
- Make sure you have maven eclipse plugin and web development environment installed on your eclipse installation.
- Next First Right click on the project and run as maven install. This will download all the libraries required to run the project. This operation may take some time depending upon internet connection.
- Second Right click on the project and select run on server. This will prompt you to select which server you wish to run the project on. Please select Apache Tomcat. This will start the project on the server.
- Use POSTMAN (Google Chrome app) to test working. The url is http://localhost:8080/ctakes/rest/ctakes/hello?text=FirstPrescription&language=language
- How to add new Drugs to cTAKES database
Run the script CreateLuceneIndexForExampleDrugs.java available on github repo. It takes in the input drugs.csv file and outputs the lucene index in drug_index directory. This folder can be then used as the drugs database input to the cTAKES.
Doctor module backend
1.Dataset
doctorModule.sql: Contains the SQL dump of the data required for the Doctor module. The database has three columns Id, Text and Type. Id is the sequential id, Text is the name of the drug/diagnosis etc while Type indicates whether it is a drug or diagnosis or any other medical term.
2.Java Sources
a. GetInformation.java
Main class which connects to the backend server to get suggestions based on the input.
b. doctorService.java
Servlet class
3.How to run is same as Patient Backend
4.pom.xml
jersey-core, jersey - server : For providing REST API to our ctakes - java based backend
jersey - servlet : For holding the servlets on the server.
jersey - json : To handle the JSON format required to format the output.
gson: For conversion to JSON format
mysql-connector-java: Driver to connect to MySQL database.
Servers hosted on Amazon web service
a. Patient: http://ec2-54-186-181-202.us-west-2.compute.amazonaws.com:8080/ctakes2/rest/ctakes/hello?text=paracetamol BID AC 10d PO, aspirin BD&language=english (example query)
b. Doctor: http://ec2-54-186-181-202.us-west-2.compute.amazonaws.com:8080/Doctor-Backend/rest/doctor/hello?text=hep (example query)
Link to Code
Screenshots